Webinar
Immuno-oncology: look beneath the surface with multiplex sequential immunofluorescence (seqIF™)
Posted on:
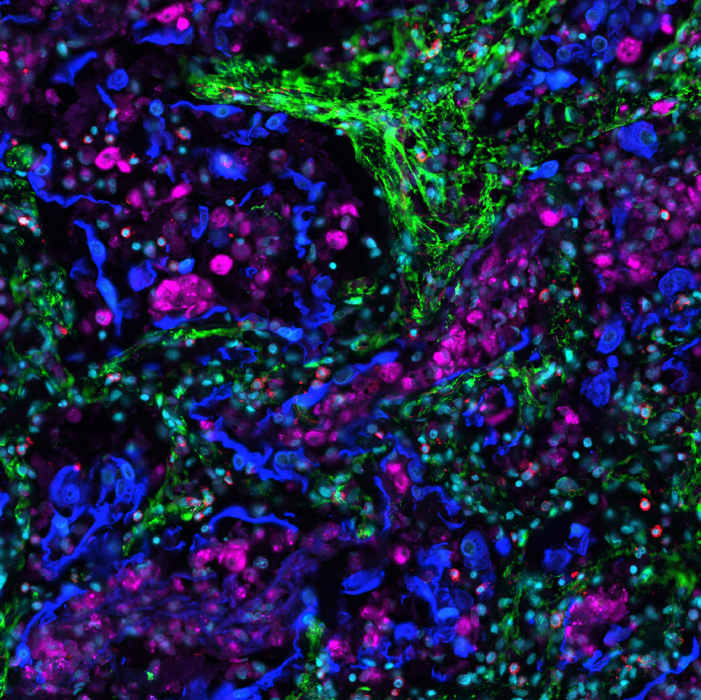
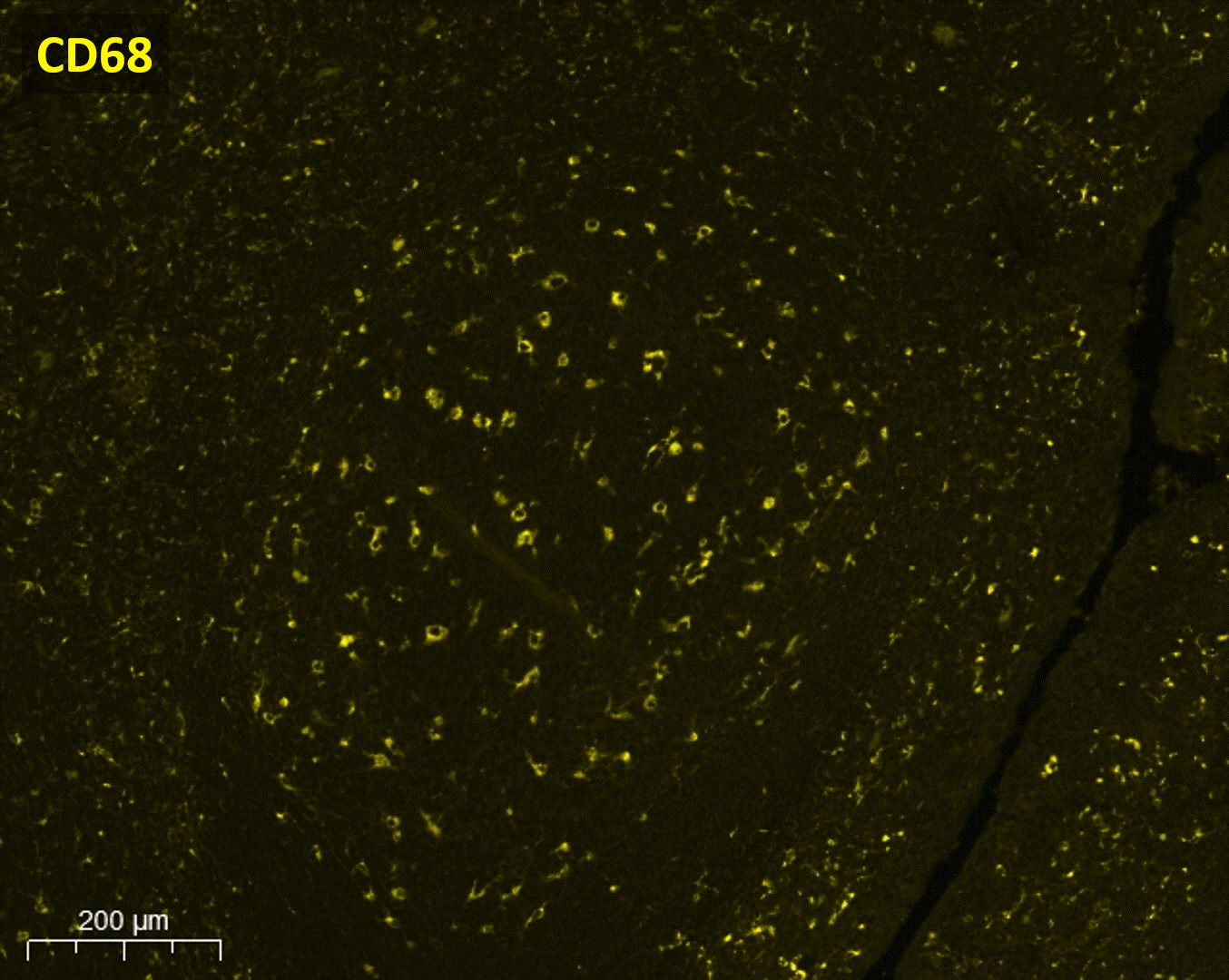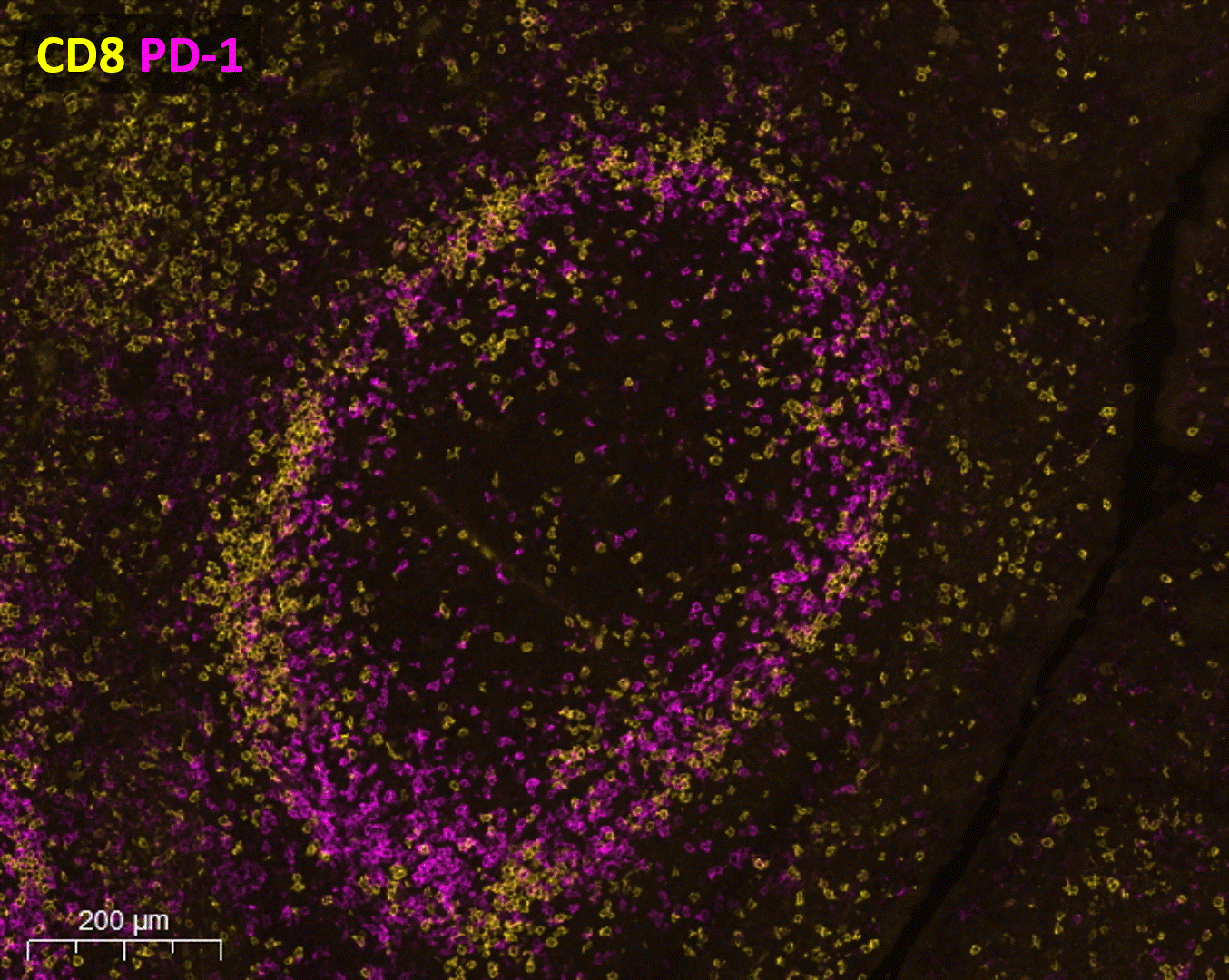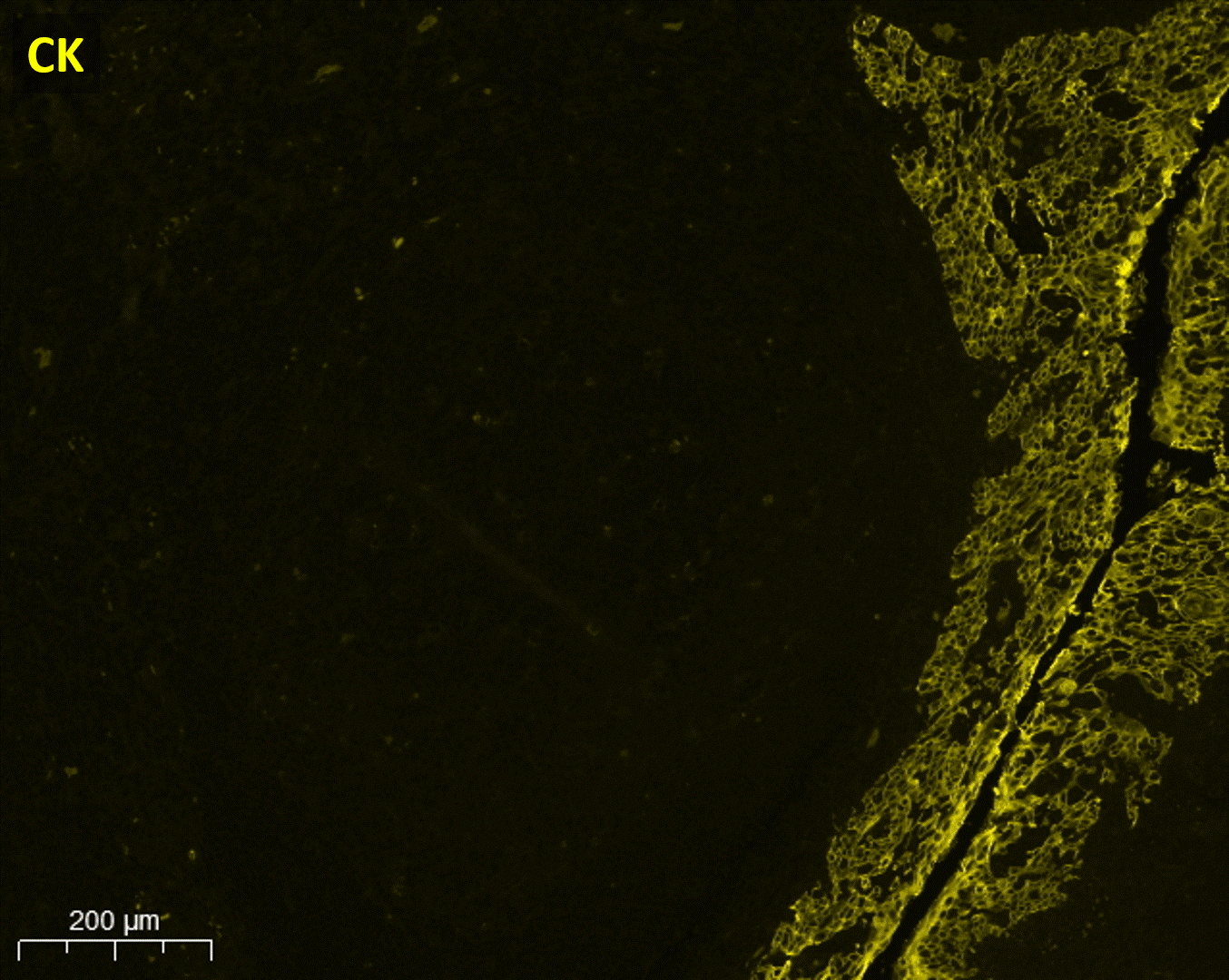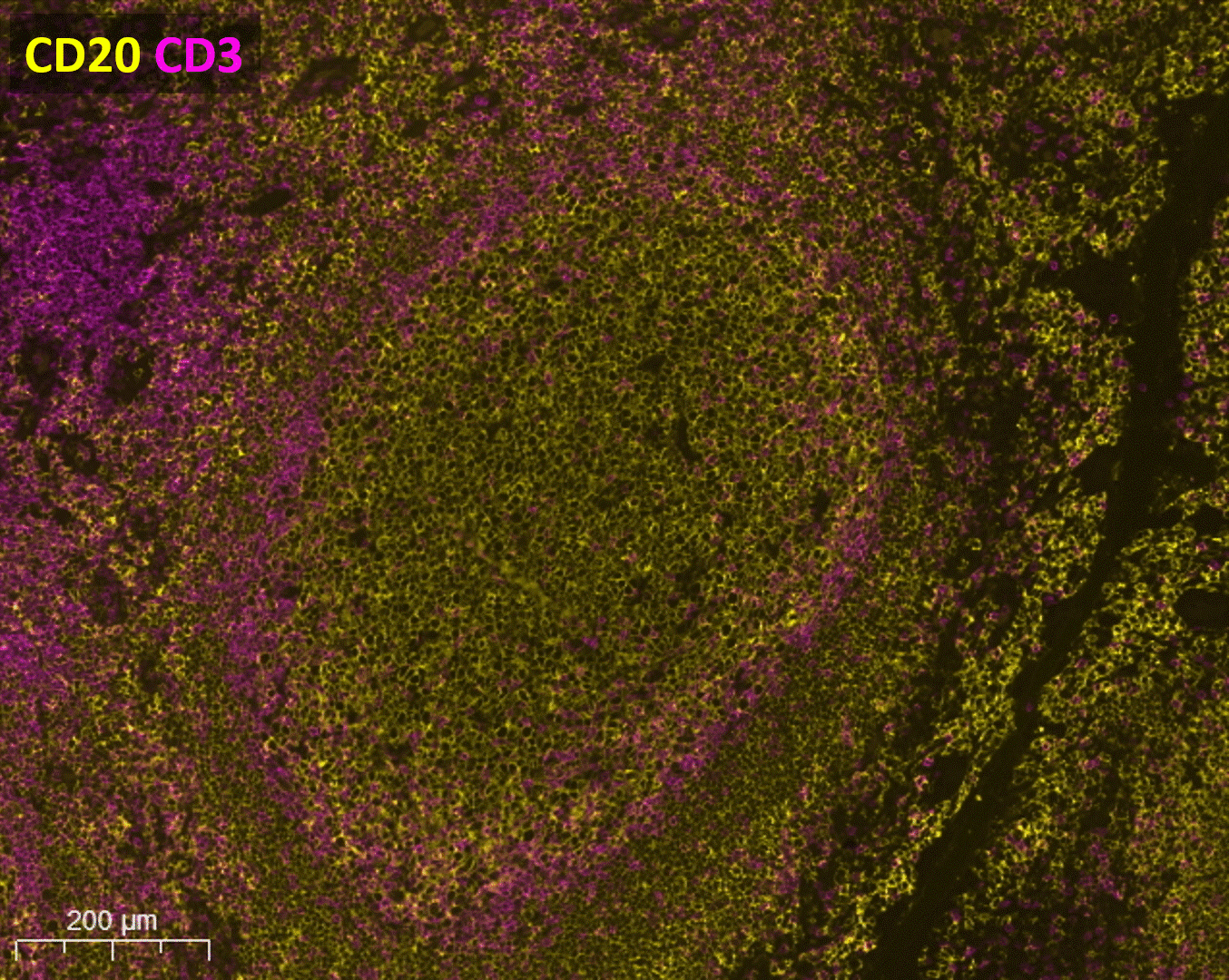
The rise of spatial biology enhances the study of the tumor microenvironment (TME) in immuno-oncology
What proteins? Where? When? – are the major questions that every scientist asks to generate a comprehensive picture of protein expression. The study of the spatial distribution of cells and their microenvironment is known as spatial biology, growing field of interest in immuno-oncology research. The interactions between various cells within the tumor microenvironment (TME) are important to investigate from a molecular standpoint. Spatial biology allows researchers to examine the distance between cells, decipher expression of molecules and generate cell density metrics. As highlighted by Dr. Paolo Ascierto, Director of the Department of Melanoma, Cancer Immunotherapy and Development Therapeutics at the National Tumor Institute in Naples, “for immuno-oncologists such knowledge is crucial to understanding the process that underlies the immune response to treatment”1.
Intensive research is currently being conducted by scientific community to identify predictors of response to anti-PD-1/PD-L1 (anti–programmed cell death 1/programmed cell death ligand 1) immunotherapy helping to match patients to the right treatment. Such discoveries also drive the target identification and development of new drugs.
What are the common challenges faced when starting off in spatial biology?
In recent years, techniques used in spatial biology have advanced substantially. Particularly spatial biology proteomics with visualization of multiple markers on the same tissue has raised considerable interest. However, for those just starting in this field, this can present several hurdles, including:
- Extensive hands-on time due to lengthy and complex protocol requirements that need substantial optimization and numerous manual steps.
- Tissue morphology degradation due to long exposure times to harsh protocol conditions.
- Lack of flexibility due to the limited offer of proprietary antibody chemistries.
- Technical limitations on the number of antigens that can be visualized at the same time depending on the chosen chemistry or the number of microscope channels.
- Limited sample availability.
Which techniques should you try first to ease adoption?
All the above-mentioned hurdles can seem daunting at first but there are some methods that are easier than others.
Researchers have a wide range of tools available to study spatial biology. Some of the existing approaches for multiple biomarker detection are the following:
- Tyramide Signal Amplification (TSA), where reagents are incubated sequentially but visualized simultaneously.
- Antibody mixes, for lower-plex stainings, where antibodies are incubated together for simultaneous imaging.
- Antibody conjugates, which involve tagging antibodies for independent detection with oligonucleotides, metal tags or breakable fluorophores.
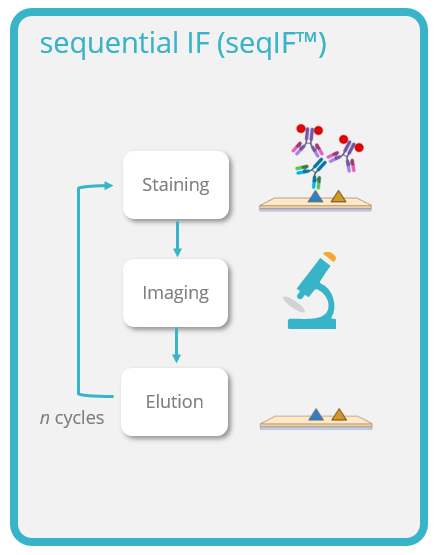
- Sequential immunofluorescence (seqIF™) which relies on standard antibodies that are incubated and eluted in a sequential manner.
Thus, scientists need to evaluate several factors to choose the best technology that supports their experimental objectives.
Related Articles
The Spatial Biology Week™ 2022: accelerating discoveries through a spatial lens
Posted on 22 Nov 2022
Read PostHORIZON™ drives insights into the glioblastoma tumor microenvironment
Posted on 27 Aug 2025
Read Post