Poster
Putting the pieces together: the spatial evolution of cancer research
Posted on:
Acknowledging and integrating the inherent heterogeneity of cancers is essential for the advancement of cancer research and the personalization of treatment modalities. The diverse functional and structural characteristics of cancers, which vary significantly across patients, present a formidable challenge in oncology, impacting treatment efficacy and prognosis. Cancer research has journeyed through an extensive and fascinating history, evolving from the high-level understanding of tumors to the sophisticated, detailed analyses we conduct today.
Evolving perspectives in cancer research
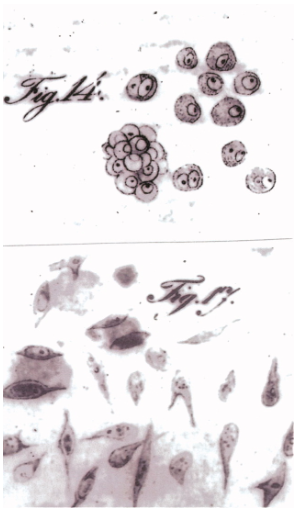
In the early 19th century, Johannes Peter Müller, a pioneering German scientist, laid the groundwork for cancer research through his initial descriptions of cancer cells1 (Figure 1). Müller’s seminal contributions marked the beginning of an era of significant breakthroughs, progressively demystifying the complexities of cancer.
In the 20th century, a significant development in cancer research was the introduction of immunohistochemistry (IHC). This technique helped identify particular tissue features and protein targets within tumor cells, transforming the way cancer is studied. IHC paved the way for more advanced characterizations of cancer by providing crucial insights into the cellular changes that cause cancer development and progression. The introduction of multiplex immunofluorescence (mIF) met the demand for advanced analytical methods by enabling the simultaneous detection of multiple markers on a single tissue section. This advancement significantly enhanced the ability to analyze the molecular complexities of the tumor microenvironment (TME). The revolution in single-cell technologies represents the latest frontier in cancer research, offering even more granular insights into cancer heterogeneity2. By enabling precise analysis at the individual cell level, these technologies are refining our understanding of the TME’s dynamics, further empowering researchers to identify novel avenues for intervention.
Grasping the spatial dynamics of the TME with integrated RNA and protein profiling
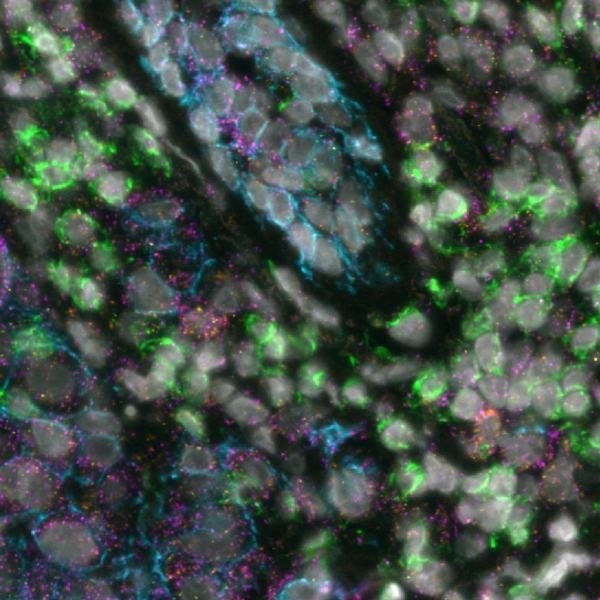
Today, spatial multiomics, which involves the simultaneous analysis of different types of molecules within their native spatial context, is revolutionizing cancer research. This technique enables the exploration of tumor heterogeneity at a molecular level, all while preserving the spatial context crucial for a comprehensive understanding of tumor biology. By delineating the molecular profiles of both cancerous and non-cancerous cells within the TME and tracking how these profiles vary across different spatial regions of the tumor, researchers can unearth the underlying mechanisms that drive tumor growth, metastasis, and resistance to treatments.
A recent study presented at the Advances in Genome Biology and Technology (AGBT) Society General Meeting 2024 highlighted a novel same-section, fully automated spatial multiomics approach on the COMET™ platform for comprehensive TME characterization. The workflow combines the RNAscope™ HiPlex Pro Assay3 with sequential immunofluorescence (seqIF™)4. Utilizing an optimized target retrieval protocol and protease-free tissue pre-treatment, this fully automated multiomics workflow supports the concurrent detection of desired RNA and protein targets (Figure 2).