Infographic
Immuno-oncology: using spatial biology to advance biomarker discovery and personalized medicine
Posted on:

Spatial analysis of protein markers dissects the heterogeneity of tumors for personalized medicine applications. Download the infographic to discover more.
Personalized medicine goes beyond diagnosis. It tailors treatments for each patient, improving patient outcomes and quality of care. Although the field has been gaining momentum in recent years, it is far from a new idea. Hippocrates’ works are where we may find the roots of our knowledge of customized treatment1. Hippocrates discussed the uniqueness of each condition and the need to treat each patient differently with the necessity of giving “different drugs to different patients, for the sweet ones do not benefit everyone, nor do the astringent ones, nor are all the patients able to drink the same things”2. Today, personalized medicine pulls together both established and innovative concepts to make drugs better and safer.
Why is it important to spatially profile the tumor microenvironment (TME)?
Tumors differ in their structural and functional characteristics, and the responses to cancer therapy may vary greatly from one patient to another. Targeted therapies focus on specific features that are particular to a given malignancy. In recent years, immunotherapy has transformed how cancer patients are treated. The goal of immunotherapy is to manipulate an individual’s own immune system and direct the immune response in order to recognize and react against malignant cells. However, to present, immunotherapy treatments have only shown promise in a small subset of tumors and in a subgroup of patients3. There is, therefore, a need to determine beforehand which cancer patients will benefit from a given immunotherapy. In addition to this, in order to deliver more potent cancer immuno- therapies that can address a larger set of patients, research efforts are concentrating on the characterization of the tumor microenvironment (TME) to uncover cell phenotypes and discover mechanisms of resistance.

The TME is the ecosystem of heterogeneous tumor cells, stromal cells (that make up connective tissue), immune cells, and various associated tissue cells (Figure 1). Traditional methods, including RNA sequencing, have revolutionized our understanding of factors influencing tumor development and therapy resistance. However, the potential of these techniques is limited because they do not give a complete picture of the tumor biology as they omit data on cellular localization within the TME4. For a thorough analysis of the TME, multiplex imaging techniques are required. Spatial biology by multiplexed immunoassays enables the simultaneous analysis of several biomarkers while preserving their morphological context within the tissue5. This technique can be used to understand the heterogeneity of cancer at many levels because it indicates cells’ presence, position and state, the density of cells, the distance between immune and cancer cells, and presents capabilities for high dimensional data analysis.
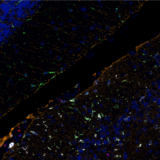
For example, multiplex immunofluorescence (mIF) panels were developed and used to characterize low-grade, early-stage endometrial carcinoma (EC) through quantitative analysis of the endometrial proteome from FFPE tissue samples6. In this study, researchers were able to use mIF analysis to demonstrate high myeloid infiltration in EC (Figure 2).

In a recent meta-analysis study, the multiplex immunohistochemistry/immuno – fluorescence (mIHC/mIF) biomarker strategy was demonstrated to be superior in predicting objective response to anti-PD-11/PD-L1 therapies than other approaches7.
The value of spatial analysis in the clinic
An increasing number of studies demonstrate that integrated knowledge obtained through mIF provides critical biological insights into cancer. For example, in the recent MITRE study, a 6-plex mIF assay to characterize the PD-1/PD-L1 axis was optimized. Highly sensitive and reproducible results were obtained with the same panel across different institutions8.
Spatially-resolved profiles are essential for identifying the molecular triggers of cellular interactions in tumors. Modern therapies rely on the accurate detection of these triggers. Therefore, improved cancer treatments will result from increased adoption of spatial technologies.
Download the infographic and discover:
- What is personalized medicine and what is the role of spatial biology in this field?
- What are predictive biomarkers?
- Why choose mIF for predictive biomarker discovery?
References:
- Steele F, Personalized medicine: something old, something new. Personalized Med. 2008; doi: 10.2217/17410541.6.1.1.
- Lepri G et al., Recent advances steer the future of systemic sclerosis toward precision medicine. Clin Rheumatol; 2020; 39(1):1-4. doi: 10.1007/s10067-019-04834-5.
- Emens L, Cancer immunotherapy: opportunities and challenges in the rapidly evolving clinical landscape. Eur J Cancer; 2017; 81:116-129. doi: 10.1016/j.ejca.2017.01.035.
- Lewis S, Spatial omics and multiplexed imaging to explore cancer biology. Methods; 2021; 18(9):997-1012. doi: 10.1038/s41592-021-01203-6.
- McGinnis LM, Ibarra-Lopez V, Rost S, et al. Clinical and research applications of multiplexed immunohistochemistry and in situ hybridization. Pathol. 2021;254(4): 405-417.
- Lopez-Janeiro A, Proteomic Analysis of Low-Grade, Early-Stage Endometrial Carcinoma Reveals New Dysregulated Pathways Associated with Cell Death and Cell Signaling. Cancers; 13(4):794. doi: 10.3390/cancers13040794.
- Lu S, Comparison of Biomarker Modalities for Predicting Response to PD-1/PD-L1 Checkpoint Blockade: A Systematic Review and Meta-analysis. JAMA Oncol; 5(8):1195-1204. doi: 10.1001/jamaoncol.2019.1549.
- Taube J, Multi-institutional TSA-amplified Multiplexed Immunofluorescence Reproducibility Evaluation (MITRE) Study. J Immunother Cancer. 9(7):e002197. doi: 10.1136/jitc-2020-002197.