Infographic
Spatial biology applications: obtain insights into biological processes with spatial proteomics
Posted on:
Are you getting started in spatial biology and wondering if this is the right solution for your research? Discover our infographic and learn more.
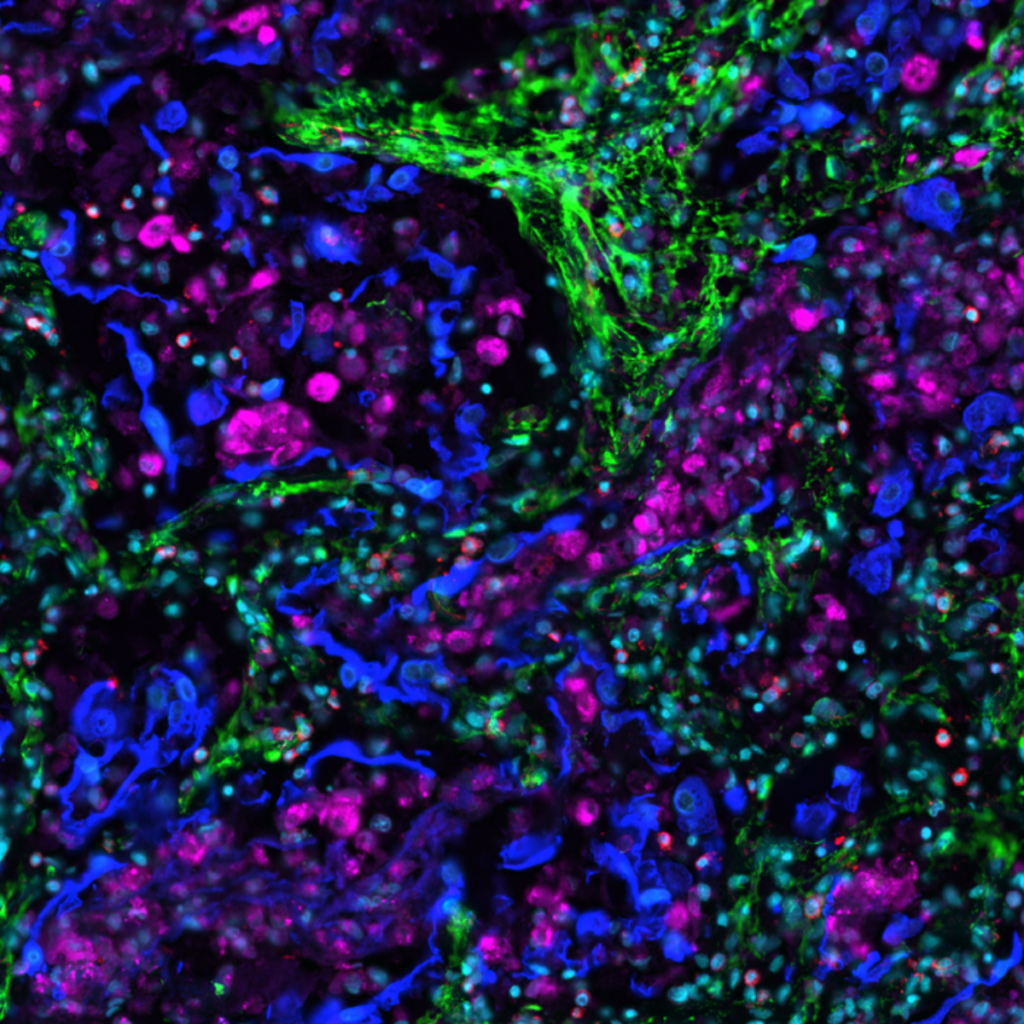
Spatial biology allows researchers to observe and measure cellular architecture. It facilitates understanding of the cellular function and cell-to-cell interactions. One of the most common subcategories of spatial biology is spatial proteomics. It entails a variety of unique approaches for obtaining information on different cell types and their dynamics.
Spatial biology enables a wide range of applications
Thorough knowledge of biological functions requires an understanding of the spatial context. Therefore, the findings that are made possible by spatial biology have the potential to transform our knowledge in many fields with key areas including:
Cancer research: In recent years, cancer treatment has been transformed by immunotherapy and patients have demonstrated encouraging results in several cancer types. The spatial approach allows the identification of specific and predictive biomarkers of response to therapy to match patients to the right treatment. In addition, it facilitates clinical oncologists to identify non-responders earlier and make better decisions.
Spatial biology tools also facilitate discoveries of new therapeutic targets in oncology to meet unmet clinical needs. For example, brain tumors are characterized by the immunosuppressive tumor microenvironment (TME)1 and high myeloid cell content2. Therefore, many patients with brain metastasis (BrM) are unable to profit from the development of immunotherapies and must rely on standard-of-care medicines with low success rates. Protein-based spatial biology allows for the characterization of neural tissue in detail. This knowledge will provide insights into therapy resistance and will aid the development of effective medicine.