Poster
Hyperplex analysis on delicate frozen samples: now possible with high-quality results
Posted on:
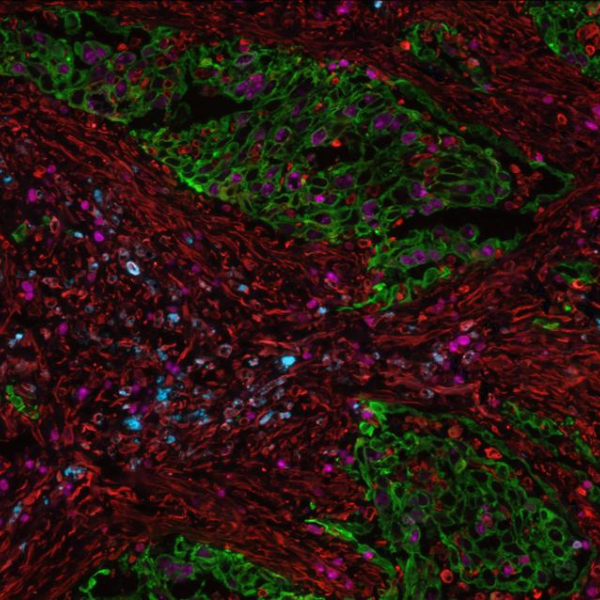
The tumor microenvironment (TME) components significantly impact cancer’s development, growth, metastasis, and the degree to which it responds or resists immunotherapies1, 2. Researchers have been widely using multiplex immunofluorescence (mIF) in recent years because it allows for a multi-layered depiction of the TME. There is a growing need for versatile, high-throughput hyperplex analysis platforms, as translational research studies increase the demand for spatial biomarker analyses3, notwithstanding the possible limited availability and the fragility of the tissue samples. In particular, there is a growing interest from researchers that are profiling TME in analyzing simultaneously multiple biomarkers on frozen sections. The existing manual, error-prone, and time-consuming procedures constrain the usage of this application. Additionally, manual protocols as well as some automated platforms often employ methods that can harm tissue morphology, such as long reagent incubations, high-temperature cycles, or photobleaching steps, which can compromise results.
Discerning protein expression in fragile frozen tissue sections using sequential immunofluorescence (seqIF™)
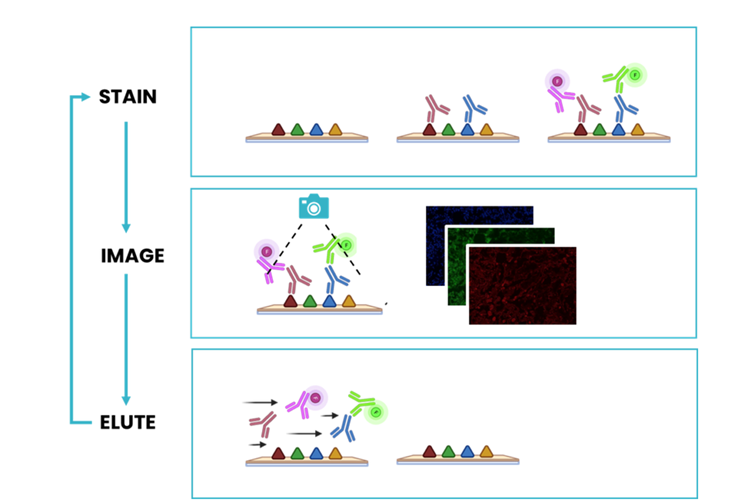
Frozen sections have the benefit of preserving the protein structure, including native antigens in each tissue. They are substantially more delicate than FFPE samples and cannot withstand stringent antigen retrieval procedures. In addition, some tissue types are particularly delicate and require the use of gentle tissue staining protocols in order to preserve tissue morphology. The revolutionary microfluidic-assisted seqIF™ method is compatible with a wide range of tissue types and allows for a comprehensive analysis of delicate samples. It consists of subsequent cycles of mIF staining, imaging, and elution, where two biomarkers are detected simultaneously in every cycle (Figure 1). The COMET™ platform performs fully automated seqIF™ assays with the detection of up to 40 biomarkers, in 20 cycles, on a single tissue slide without user intervention. Thanks to the patented microfluidic chip technology, all conditions are precisely controlled, enabling seqIF™ experiments on COMET™ with exceptional reproducibility, signal uniformity, and highly preserved tissue morphology.
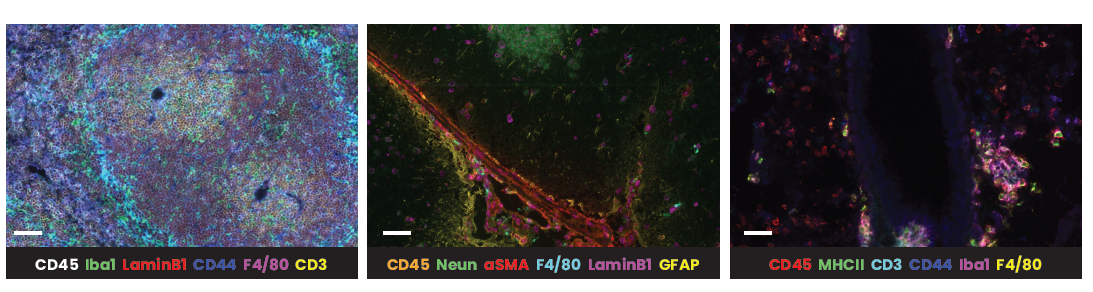
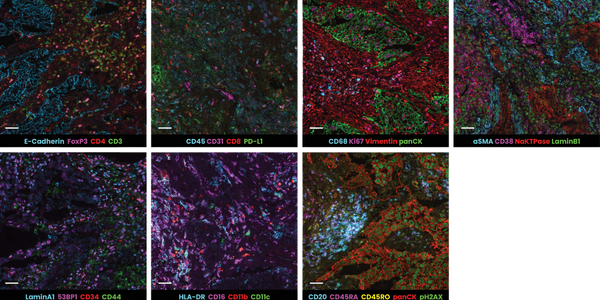
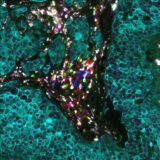
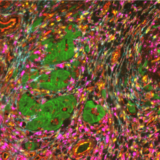
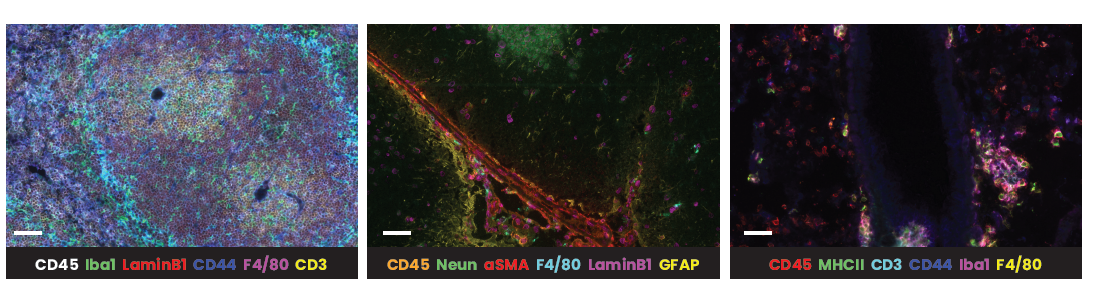
In the example below, murine spleen, brain, and lung tissues were analyzed with multiplex panels demonstrating accurate detection of the specified biomarkers to assess the compatibility of delicate murine samples with the seqIF™ protocol on COMET™ (Figure 2).
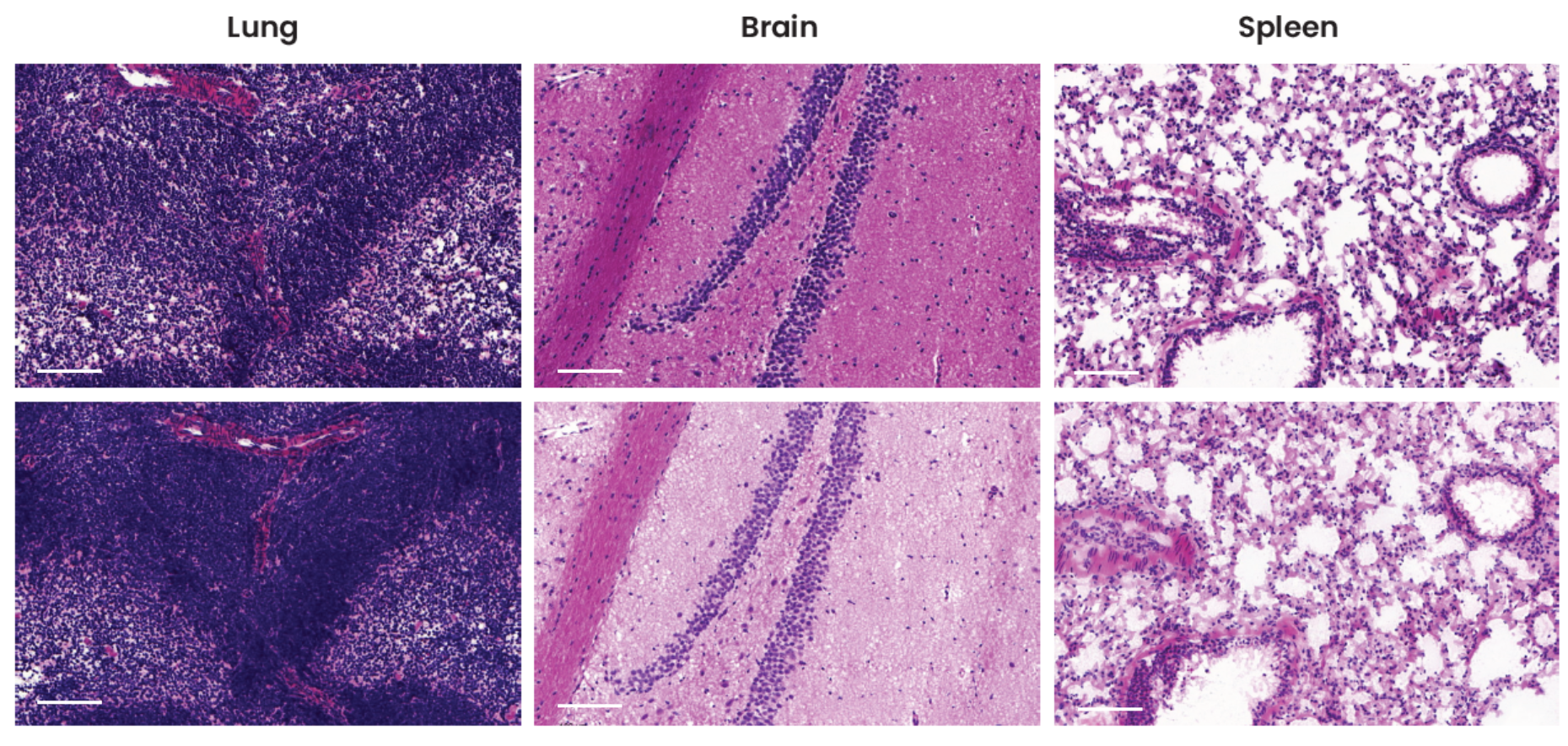
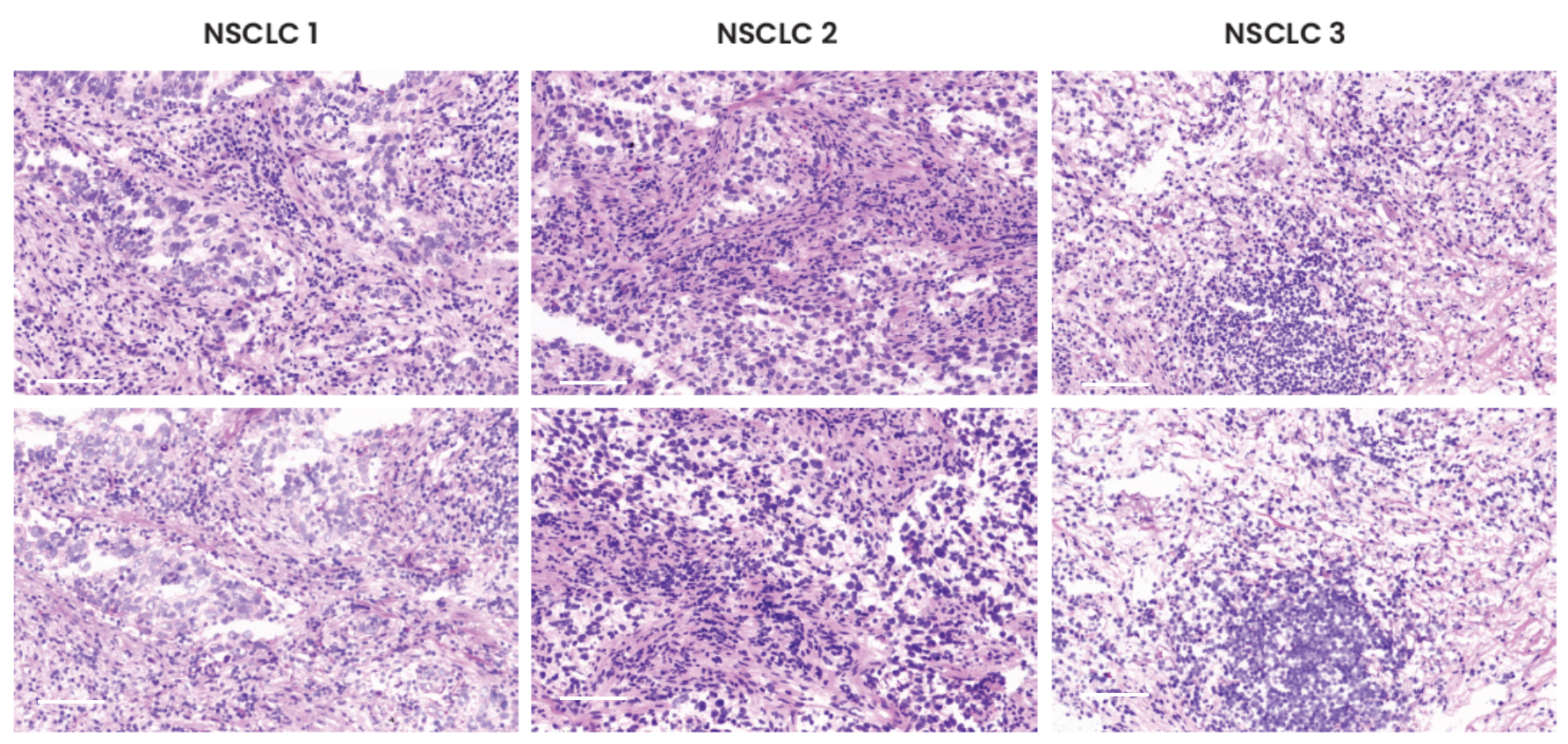
Pathologists have used hematoxylin and eosin (H&E) staining to visualize multiple tissue components in microscopic examinations of biopsies and surgical samples for more than a hundred years4. In addition to mIF, researchers can benefit from using this traditional method on the same slide in order to analyze the tissue architecture and morphological structure. The preservation of tissue morphology is key to performing any additional downstream applications after a multiplex assay, such as H&E or even chromogenic IHC. A study was performed to assess tissue morphology preservation after a seqIF™ protocol. The specimens were further stained with H&E after undergoing a 20-cycle protocol on COMET™, equivalent to a 40-plex analysis. As demonstrated by H&E staining run after seqIF™, as compared to freshly processed control slides, the tissue morphology for all three tissue types was well preserved (Figure 3).
Related Articles
Immunosuppressive brain tumor microenvironment: a barrier to overcome
Posted on 01 Jun 2023
Read Post